Polymerase Chain Reaction
The polymerase chain reaction (PCR) is used to make millions of copies of a section of deoxyribonucleic acid ( DNA ). Until the 1980s obtaining numerous copies of a section of DNA took one to two weeks and required isolation of the DNA, cloning the DNA into a viral or plasmid vector, growing the cloned DNA using living host cells, usually bacteria, and finally isolating the DNA again. With PCR a scientist can produce thirty million copies of a DNA section in a test tube overnight. In a series of early articles, PCR's inventor, Kary Mullis, described just how valuable he believed this tool would be. He could not have been more correct: PCR is now a mainstay of molecular biology. In 1993 Mullis received the Nobel Prize in chemistry in recognition of his discovery.
How PCR Works
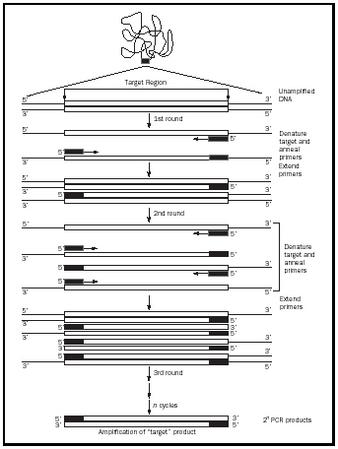
PCR repeats the synthesis of a DNA segment through twenty-five to thirty cycles. The power of the technique resides in its ability to copy the products of each previous cycle along with the original sample. This results in exponential growth in the amount of product, as illustrated in Figure 1. The first stage or round of the process only involves copying the original sample. First, the DNA is separated into its two strands, and each strand acts as an instruction directing a new segment. The two products each contain one original strand and one new one. In the second round of the process, both of the DNA molecules manufactured in stage one are separated and each of the four strands is copied, producing four double-stranded products. The number of DNA segments doubles during each round. Twenty-five rounds can create millions of new segments from each original DNA.
To separate the double-stranded DNA products, the reaction is heated to near boiling, 95°C, for a few minutes. Originally, the catalyst, DNA polymerase, was killed by this treatment and manually replaced after each round. In 1988 Randall Saiki and coworkers dramatically improved the original concept of PCR by using a heat-stable catalyst, taq polymerase, isolated from the bacteria, Thermis aquaticus , found in hot springs. This catalyst survives the high temperature step and can start the next round of synthesis when cooled to the start temperature.
The specificity of PCR derives from the primers. Primers are short pieces of single-stranded DNA that are the starting point of the new DNA strands. The primers recognize the target DNA segment in the sample and
bind to the target. Taq polymerase uses the primer as the beginning of the new product strand. Taq polymerase cannot synthesize DNA without a primer at the beginning, so the primers and not the catalyst determine what segment of the sample is copied.
Applications of PCR
Originally, PCR was employed to produce usable amounts of small- and medium-sized DNA segments. Many other important techniques quickly developed. PCR is used to identify bacteria and virus infections. It is not only very sensitive but is also selective enough to distinguish closely related strains. For example, this procedure has been adapted to detect genetically modified crops to help ensure they are utilized only in approved ways.
A widely known application of PCR is DNA fingerprinting. Certain regions of the human chromosome have short sections that are repeated up to several hundred times. Different individuals have different numbers of repeats in each section. If several sections are tested, a unique pattern is observed. Initially, these regions were cut out of sample DNA and tested for size. PCR allows the variable regions to be copied millions of times, greatly increasing the sensitivity and speed of the technique. Because of the technique's sensitivity, extreme care must be taken to avoid any contamination of the samples. Forensic scientists continue to make even greater use of this method.
One of the latest applications of PCR is the DNA microarray or gene chip. It allows medical personnel to test a cancer cell before and after chemotherapy with different drugs. If two patterns look the same, the two drugs administered are likely working on the same pathway and may not be as useful as treatment with two drugs that work by different pathways.
SEE ALSO Deoxyribonucleic Acid (DNA) ; Forensic Chemistry .
David Speckhard
Bibliography
Erlich, Henry (1995). "PCR Technology." In Molecular Biology and Biotechnology , ed. Robert Meyers. New York: Wiley-VCH.
Guyer, R., and Koshland, Daniel (1989). "The Molecule of the Year." Science 246: 1543.
Joyce, G. (1992). "Directed Molecular Evolution." Scientific American 267: 90.
Mullis, Kary (1990). "The Unusual Origin of the Polymerase Chain Reaction." Scientific American 262: 56.
Comment about this article, ask questions, or add new information about this topic: